How can you choose a single isoform to best represent a gene?
I asked this question a few weeks ago. There are some situations in bioinformatics when you want to look at all genes from one or more organisms, but where you want to only have one representative of a gene. This is not always a straightforward task, and I previously discussed how many people opt for overly simplistic methods such as 'choosing the longest isoform'.
Since my blog post I had some feedback and have also come across a relevant paper which I would like to share here.
Feedback
Sara G said:
We use epigenetic data from ENCODE/modENCODE/Epigenetic roadmap or our own data to identify the transcription start site that has the most evidence of transcriptional activity.
From my experience with using data from fly, worm, and Arabidopsis I have often seen many genes which have multiple isoforms that all share the same transcription start site (and differ in downstream exon/intron structure).
Michael Paulini commented:
What you can easily do (if you got the data) is just pick the isoform with the highest FPKM value per gene.
The important caveat that Michael mentions is that you might not have the data, and if you are trying to do this as part of multiple species comparison then this becomes a much more complex task.
Richard Edwards added to the debate (I've edited his response a little bit):
My previous solution to this problem was this:
"…a dataset consisting of one protein sequence per gene was constructed from the EnsEMBL human genome…by mapping all EnsEMBL human peptides onto their genes and assessing them in the context of the external database entry used as evidence for that gene. If the external database was SwissProt and one of the peptides has the exact same sequence as the SwissProt sequence, this peptide sequence was used for that gene. In all other cases, the longest peptide (in terms of non-X amino acids) was used."
Richard also conceded that this approach "only works for well annotated proteomes".
Another approach
For Arabidopsis thaliana, I have previously used data from their exon confidence ranking system which ends up producing a 5 star rating for every transcript of a gene. This is based on various strands of evidence and can be useful for choosing between isoforms which have different star ratings (but not so helpful when all isoforms have the same rating). The TAIR database remains the only model organism database (that I know of) that have attempted to address this problem and provide a systematic way of ranking all genes. Anecdotally, I would say that not many people know about this system and you need to find the relevant files on their FTP site to make use of this information.
Hopefully someone will correct me if other databases have implemented similar schemes!
From the literature
If your research focuses on vertebrates, then this approach looks promising.
Here's the abstract (emphasis mine):
Here, we present APPRIS (http://appris.bioinfo.cnio.es), a database that houses annotations of human splice isoforms. APPRIS has been designed to provide value to manual annotations of the human genome by adding reliable protein structural and functional data and information from cross-species conservation. The visual representation of the annotations provided by APPRIS for each gene allows annotators and researchers alike to easily identify functional changes brought about by splicing events. In addition to collecting, integrating and analyzing reliable predictions of the effect of splicing events, APPRIS also selects a single reference sequence for each gene, here termed the principal isoform, based on the annotations of structure, function and conservation for each transcript. APPRIS identifies a principal isoform for 85% of the protein-coding genes in the GENCODE 7 release for ENSEMBL. Analysis of the APPRIS data shows that at least 70% of the alternative (non-principal) variants would lose important functional or structural information relative to the principal isoform.
It seems that since this paper was published, they have expanded their dataset and the online APPRIS database now has annotations for five vertebrate genomes (human, mouse, zebrafish, rat, and pig).
The future
This APPRIS approach seems promising, yet I wish there was an standardized ranking system used by all model organism databases (as well as Ensembl, UCSC Genome Browser etc.).
There are always going to be different levels of supporting evidence available in different species. E.g. not every organism is going to be able to make use of mass spec data to assess the differential usage of transcript variants. Furthermore, there are going to be tissue-specific — not to mention time-specific — patterns of differential isoform expression for some genes.
However, what I would love to see is each database using whatever information they have availble to them to assign relative 'weights' to each annotated isoform. Initially, this would focus on protein-coding genes and ideally, this scheme would entail the following:
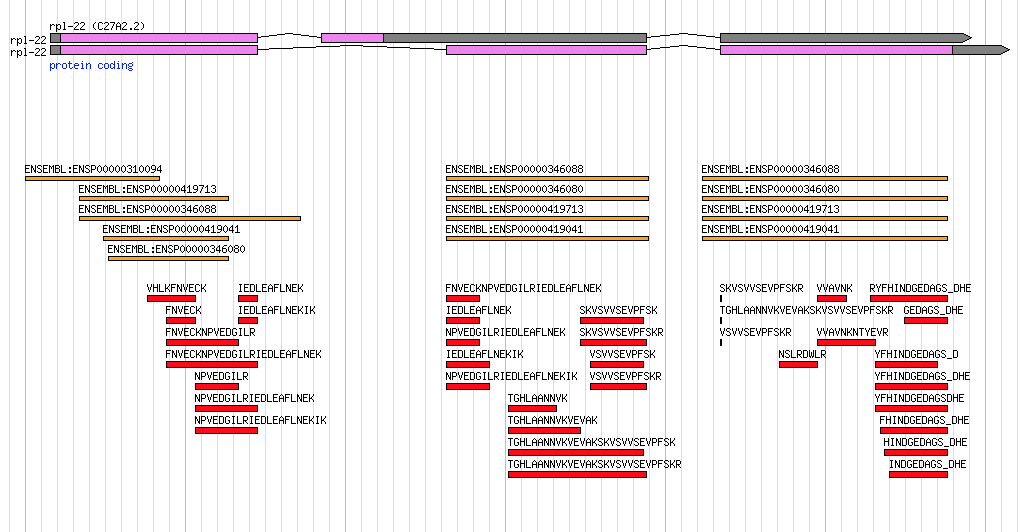
- A score for every annotated isoform of a gene where all scores sum to 100. In my favorite example of the Caenorhabditis elegans rpl-22 gene these scores would probably end up as 99 for the dominant isoform and 1 for the secondary isoform. The scores should reflect the relative likelihood that this transcript encodes a functional protein. To keep things simple, I would prefer integer values with values of zero allowed for annotated isoforms that represent pseudogenic transcripts, targets of nonsense-mediated decay etc.
- Evidence for the score. This would be something akin to the Gene Ontology Evidence codes. Each gene may have one or more codes attached to them and these would reflect the source of the experimental evidence (transcript data, mass spec, functional assay etc.)
- Spatial/temporal information. If some of the supporting evidence is from a particular cell-line, tissue, or developmental time point then it should be possible to annotate this.
- Historical information. Such scores should be expected to change over time as new evidence emerges. All transcripts should keep a time-stamped history of previous scores to allow researchers to see whether isoforms have increased or decreased in their relative probability of usage over time.
- Data to be added to GTF/GFF files: The score of each transcript should be embedded as a note in the attribution column of these files, maybe as a 'RF' score (Relative Functionality).
Can someone please make this happen? :-)